OLIGOSACCHARIDE ANALYSIS
If you find any inaccurate information, please let us know by providing your feedback here

Tóm tắt nội dung
This article is compiled based on the United States Pharmacopeia (USP) – 2025 Edition
Issued and maintained by the United States Pharmacopeial Convention (USP)
1 INTRODUCTION
Analysis of the Asparagine (Asn-) linked oligosaccharide (also known as the N-linked oligosaccharide or N-glycan) composition of selected recombinant therapeutic proteins has become necessary for product characterization or release specifications of these proteins. For information on the characterization and assessment of protein glycosylation, see Glycoprotein and Glycan Analysis—General Considerations 〈1084〉. Chapter 〈1084〉 covers general analytical strategies and also highlights criteria for selecting appropriate methods for specific analytical challenges, thereby providing the context for this chapter. This chapter focuses on the analysis of N-linked oligosaccharides that are released from glycosylated recombinant therapeutic proteins. Therefore, the chapter provides 1) a generic procedure for enzymatic release of N-glycans using peptide N-glycosidase F (PNGase F), 2) two different approaches for fluorophore labeling of the released N-linked oligosaccharides, and 3) five analytical procedures based on chromatographic and electrophoretic separation, as well as performance criteria for the separation. Alternative analytical strategies are discussed in 〈1084〉. Although the procedures in this chapter are validated, the procedures must be verified with the individual specific product (see Verification of Compendial Procedures 〈1226〉). Furthermore, validation is required when the procedure is optimized for a specific product [e.g., outside the allowed parameter limits defined in Chromatography 〈621〉, System Suitability (see Validation of Compendial Procedures 〈1225〉)]. Data analysis, quantitation, and lot release specifications, which are expected to be product specific, will be found in individual product monographs; therefore, these aspects are not covered in this chapter.
Labeling oligosaccharides with a fluorophore before analysis provides an important advantage of enhanced detection of oligosaccharides during separation, because oligosaccharides have no or low absorptivity of either ultraviolet (UV) or visible light and no significant fluorophore groups. This chapter describes two different methods for fluorophore labeling of oligosaccharides: derivatization of oligosaccharides using 2-aminobenzamide (2-AB) for liquid chromatographic separation, and labeling of oligosaccharides with 8-aminopyrene-1,3,6-trisulfonic acid (APTS) for capillary electrophoresis (CE) separation. Although electrochemical detection (pulsed amperometric detection or pulsed electrochemical detection) is also used in oligosaccharide analysis by using a strong base elution, the electrochemical detection method is less sensitive and has a higher noise-to-signal ratio. Therefore, this detection method is not addressed in this chapter.
Many different chromatography and electrophoresis methods have been developed to analyze oligosaccharides, and the procedures described in this chapter have been used successfully for this purpose. The choice of procedures discussed is meant to reflect the most commonly and broadly used approaches for supporting current recombinant glycosylated biological medicines in commerce. These analytical procedures include normal phase or hydrophilic interaction liquid chromatography (HILIC), high-performance anion exchange liquid chromatography (HPAEC), and CE. Several other chromatographic modes that separate oligosaccharides by size, shape, and polarity are also commonly used for oligosaccharide separation, but those other chromatographic modes are not discussed in this chapter. The general principles of chromatography are covered in 〈621〉, and therefore 〈621〉 should be considered within that context.
USP Reference Standards (RS) have been developed to assess system suitability for these analytical procedures:
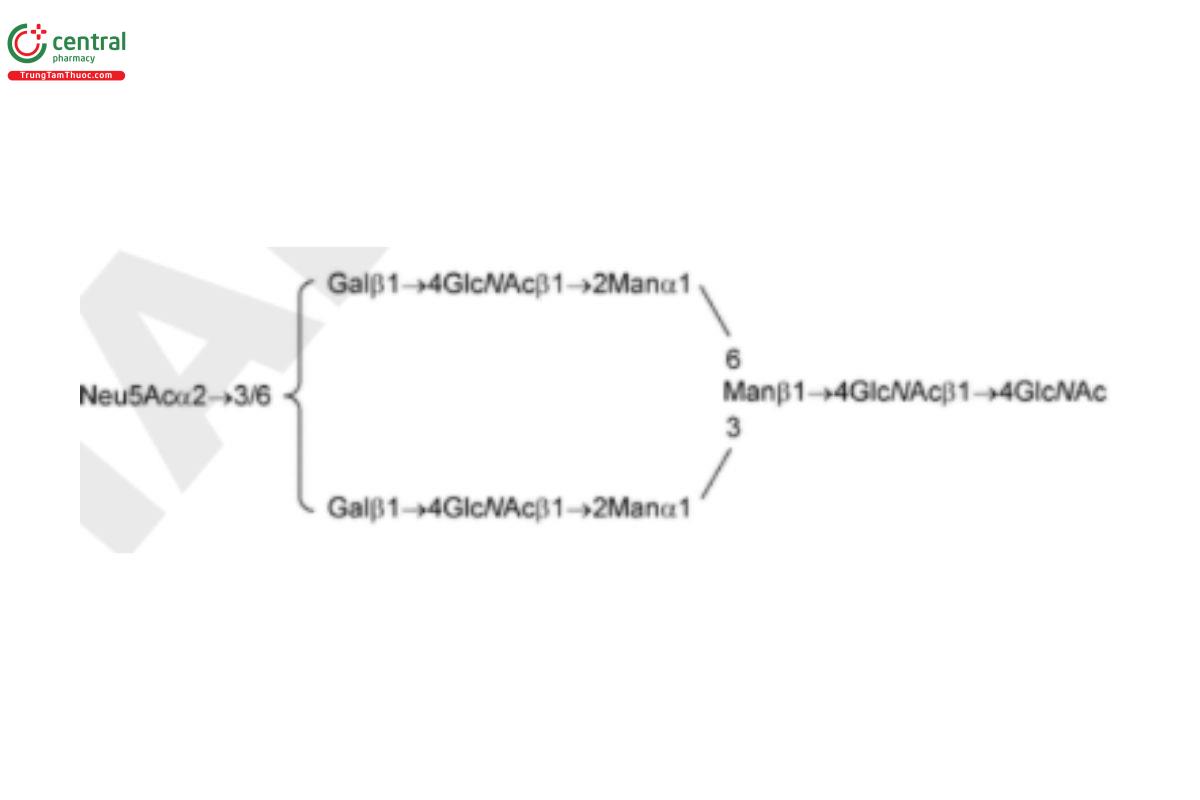
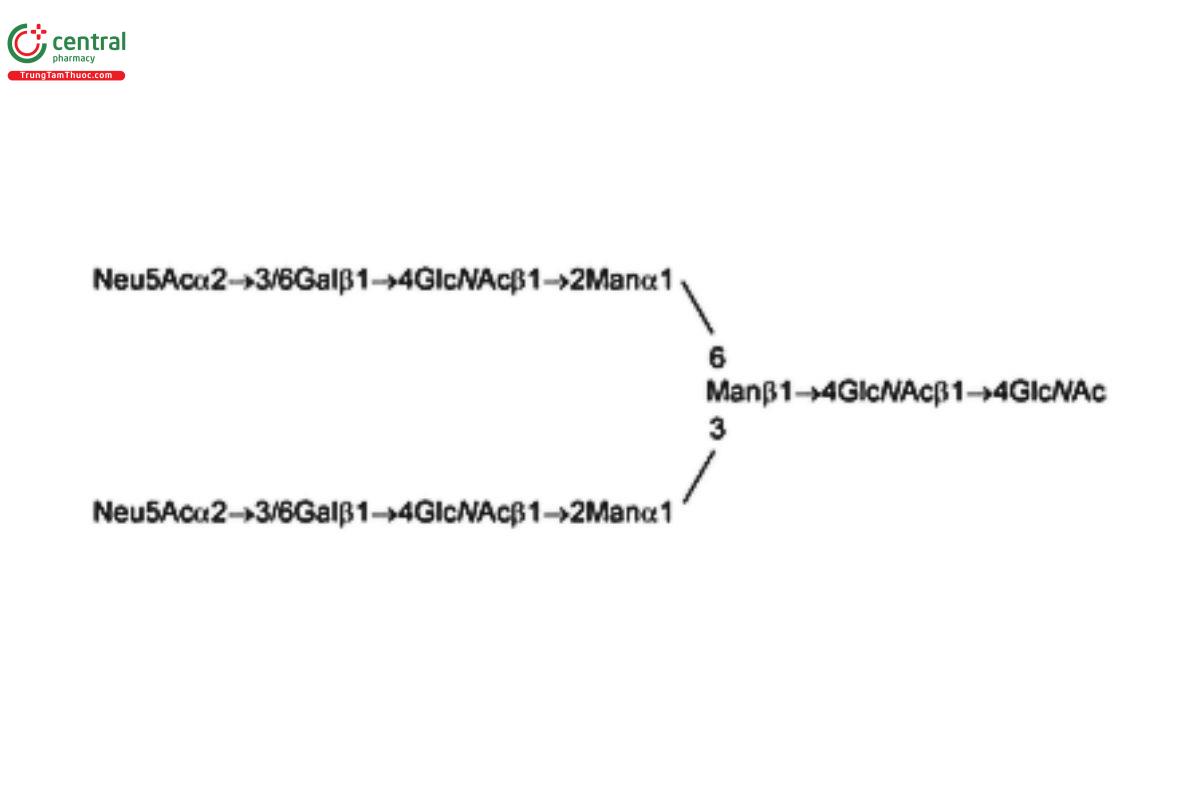
USP Oligosaccharide System Suitability Mixture A RS consists of a mixture of partially sialylated, partially galactosylated, and partially fucosylated biantennary N-linked oligosaccharides that were released from human polyclonal Immunoglobulin G (IgG) by PNGase F.
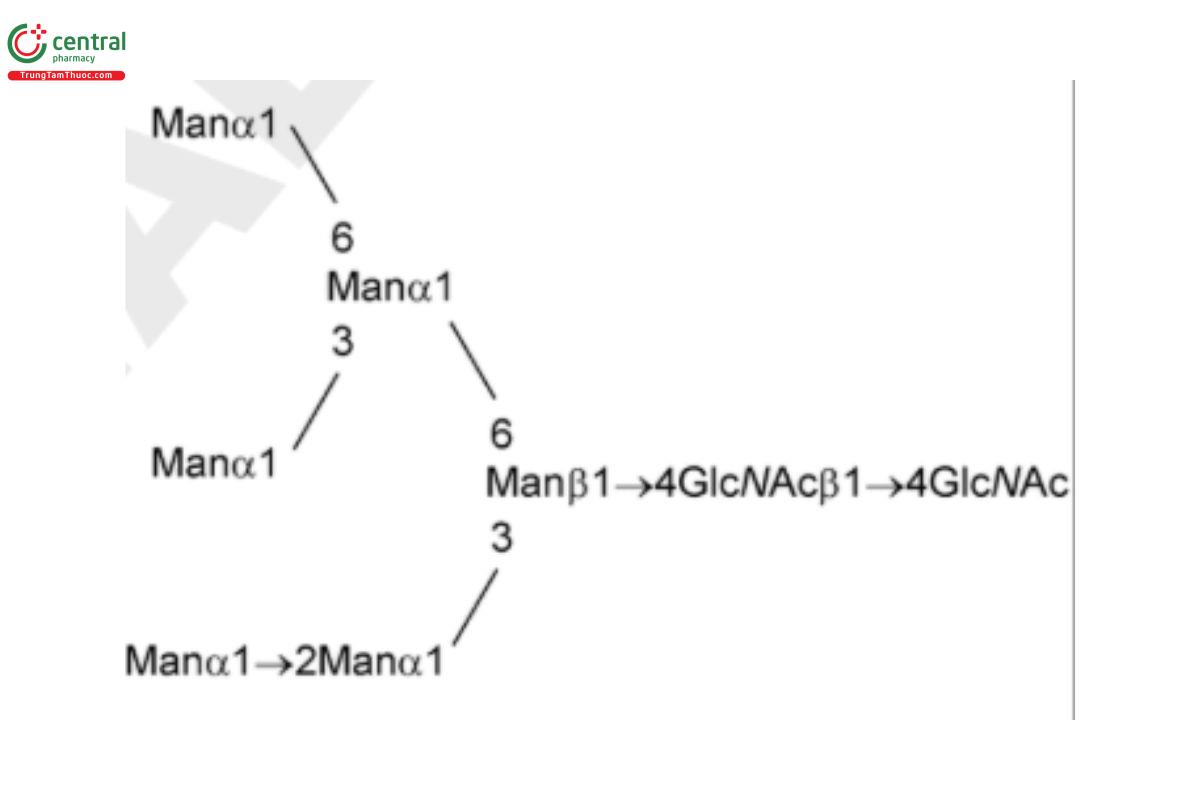
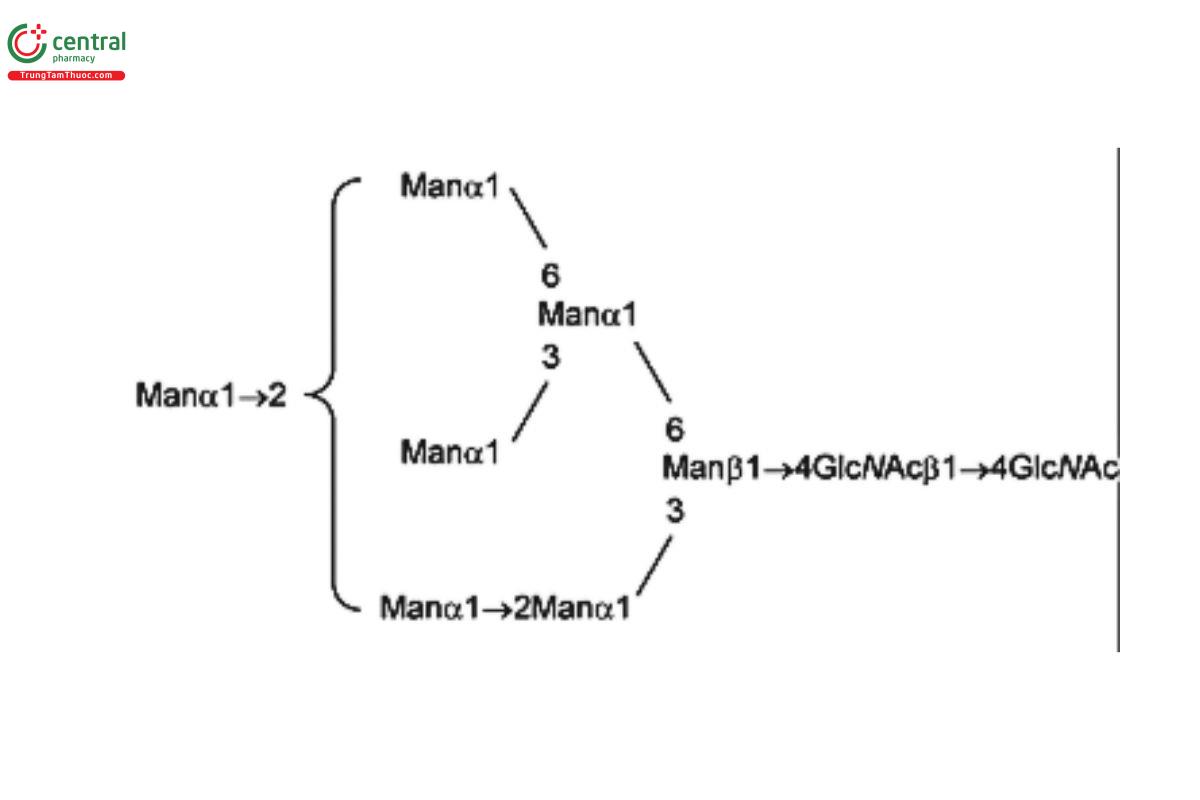
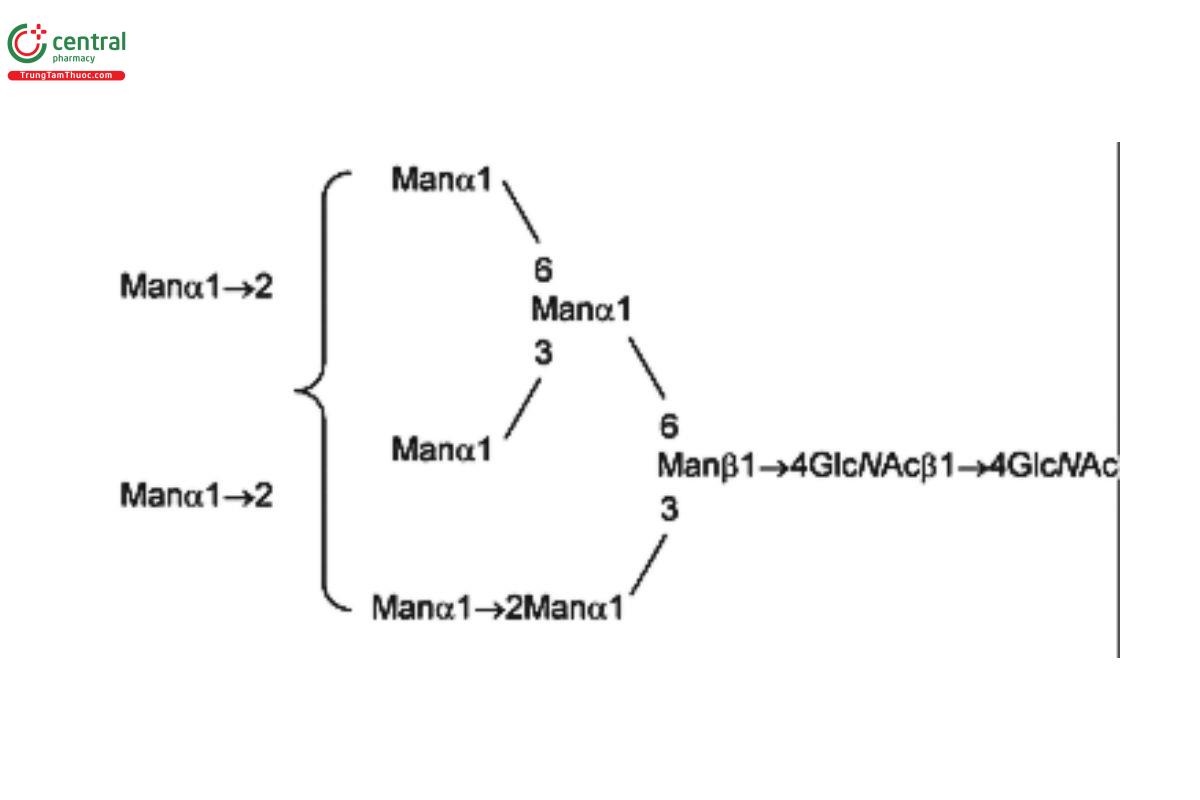
USP Oligosaccharide System Suitability Mixture B RS consists of a mixture of high-mannose N-linked oligosaccharides with traces of hybrid chains that were released from bovine ribonuclease B (RNase B) by PNGase F.
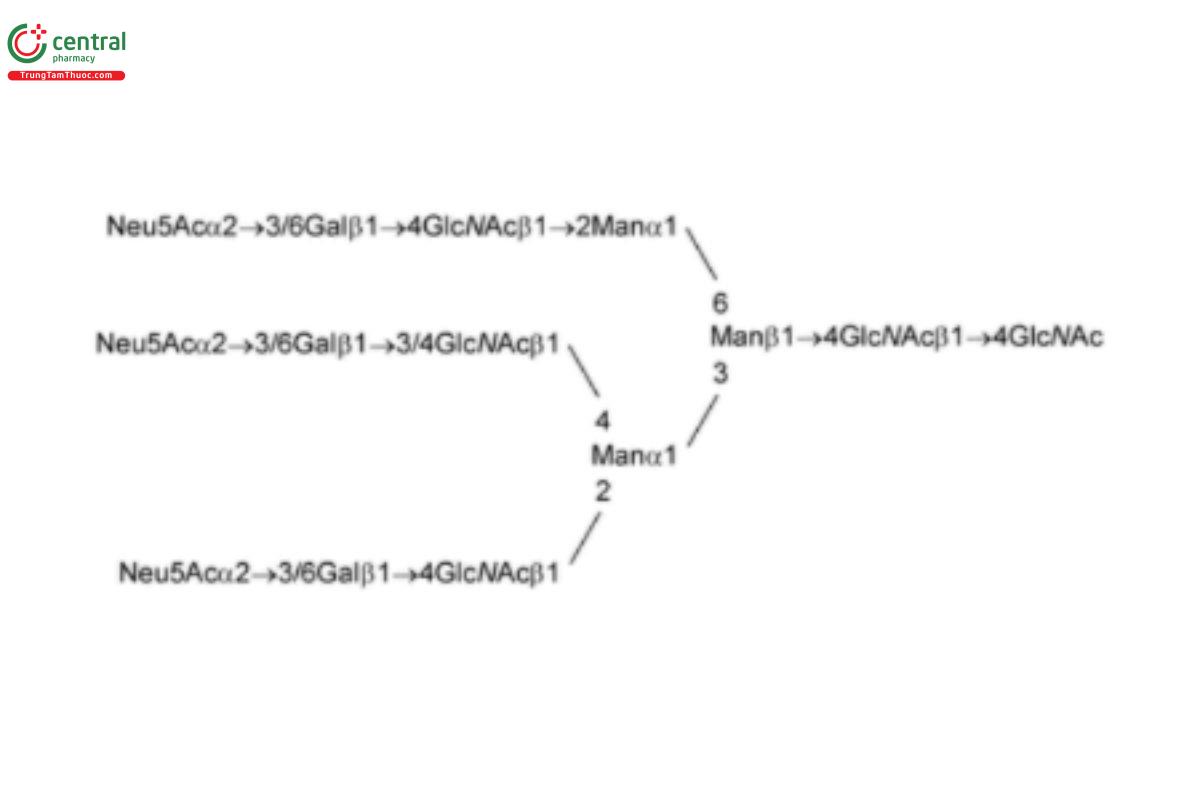
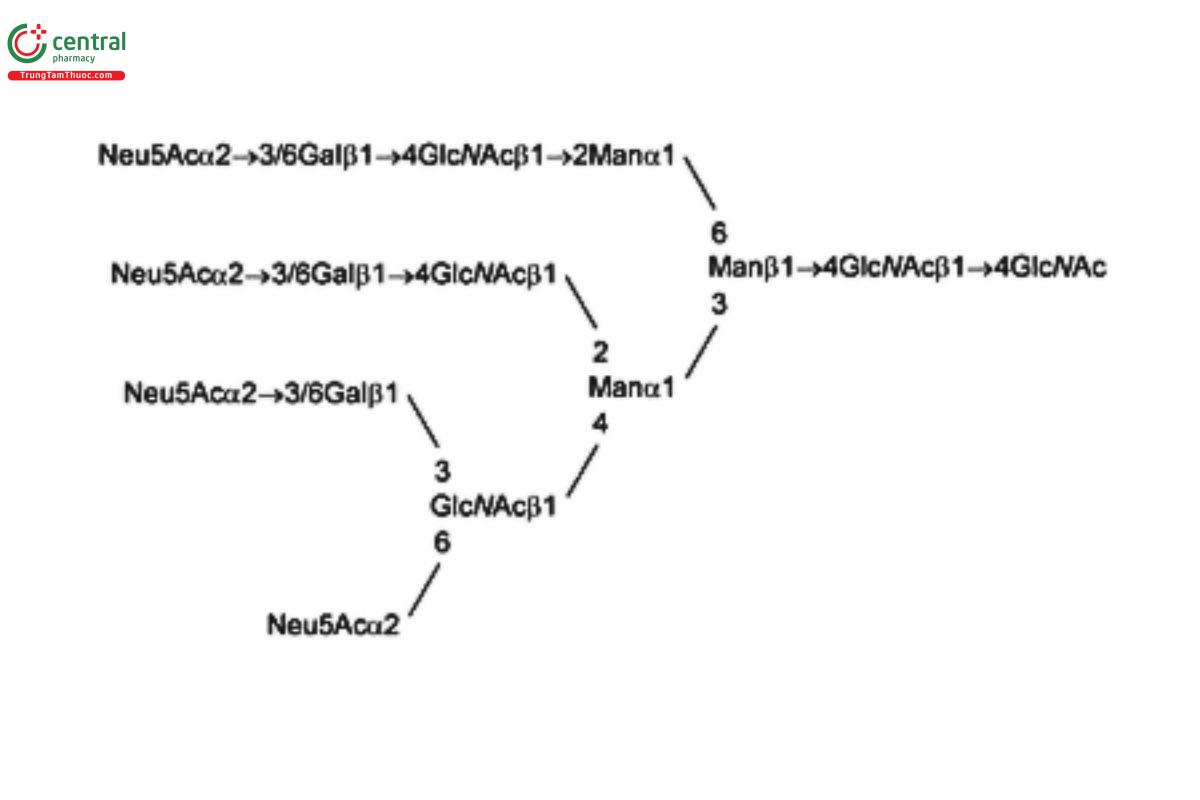
USP Oligosaccharide System Suitability Mixture C RS is a mixture of bi-, tri-, and tetra-antennary N-linked oligosaccharides with variable sialylation with or without fucose (Fuc; also containing Lex-type structures); the mixture was released from human α1 acid glycoprotein by PNGase F.
USP Oligosaccharide System Suitability Mixture D RS is a mixture of sialylated bi- and tri-antennary complex type structures with an additional sialic acid on one of the tri-antennary structures; the mixture was released from bovine fetuin by PNGase F.
2 ANALYTICAL PROCEDURES OVERVIEW
Table 1 outlines the applications and separation principles of the analytical procedures included in this chapter. The choice of the appropriate RS will depend on the expected glycan content of the product being analyzed.
| Analytical Procedures | Type of N-linked Oligosaccharides to Be Separated | Applicable USP Reference Standards |
|---|---|---|
| Normal phase liquid chromatography/HILIC procedure 1: Column: 2.0-mm × 15-cm; 3-μm packing L68 More oligosaccharide peaks compared to HILIC procedure 2 Longer mobile phase with shallow gradient compared to HILICprocedure 2 Enhanced sensitivity for smaller sample size | Relatively simple biantennary chains with no or low levels of sialylated structures, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture A RS |
| High-mannose chains, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture B RS | |
| Normal phase liquid chromatography/HILICprocedure 2: Column: 4.6-mm × 25-cm; 5-μm packing L68 Fewer oligosaccharide peaks observed compared to HILICprocedure 1 Shorter mobile phase with less shallow gradient compared to HILIC procedure 1 | Relatively simple biantennary chains with no or low levels of sialylated structures, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture A RS |
| High-mannose chains, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture B RS | |
| HPAECprocedure 1 Guard column: 4.0-mm × 5-cm; 10- μm packing L46 Column: 4.0-mm × 25-cm; 10-μm packing L46 Mobile phase: 0–75 mM sodium acetate in isocratic 0.15 N sodium hydroxide Selectivity is different from HILIC procedures 1 and 2 with respect to the size and composition of oligosaccharide | Relatively simple biantennary chains with no or low levels of sialylated structures, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture A RS |
| High-mannose chains, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture B RS | |
| HPAEC procedure 2 Guard column: 4.0-mm × 5-cm; 10- μm packing L46 Column: 4.0-mm × 25-cm; 10-μm packing L46 Mobile phase: 150–450 mM sodium acetate in isocratic 0.15 N sodium hydroxide Suitable for separation of charged N-linked oligosaccharides | Bi-, tri-, and tetra-antennary N-linked oligosaccharides with variable sialylation, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture C RS |
| Sialylated bi- and tri-antennary complex type structures with an additional sialic acid on one of the tri-antennary structure, after 2-AB labeling | USP Oligosaccharide System Suitability Mixture D RS |
3 SAMPLE PREPARATION
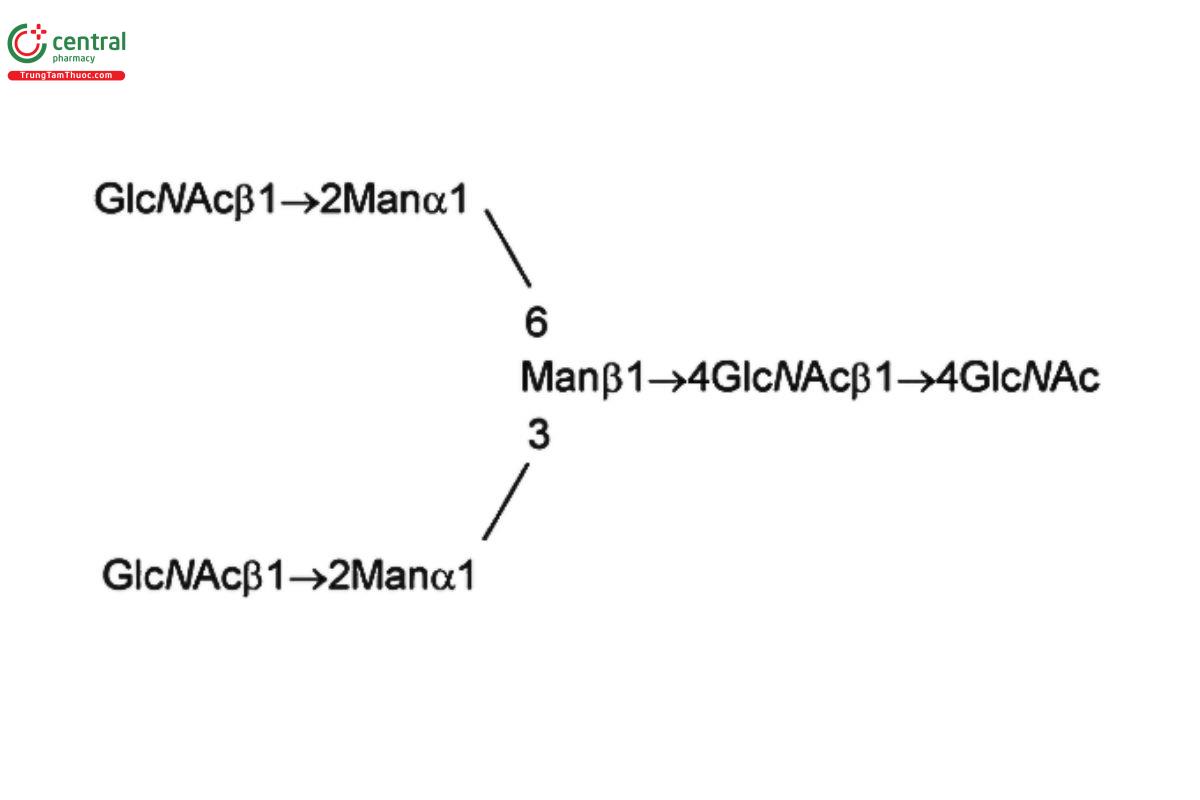
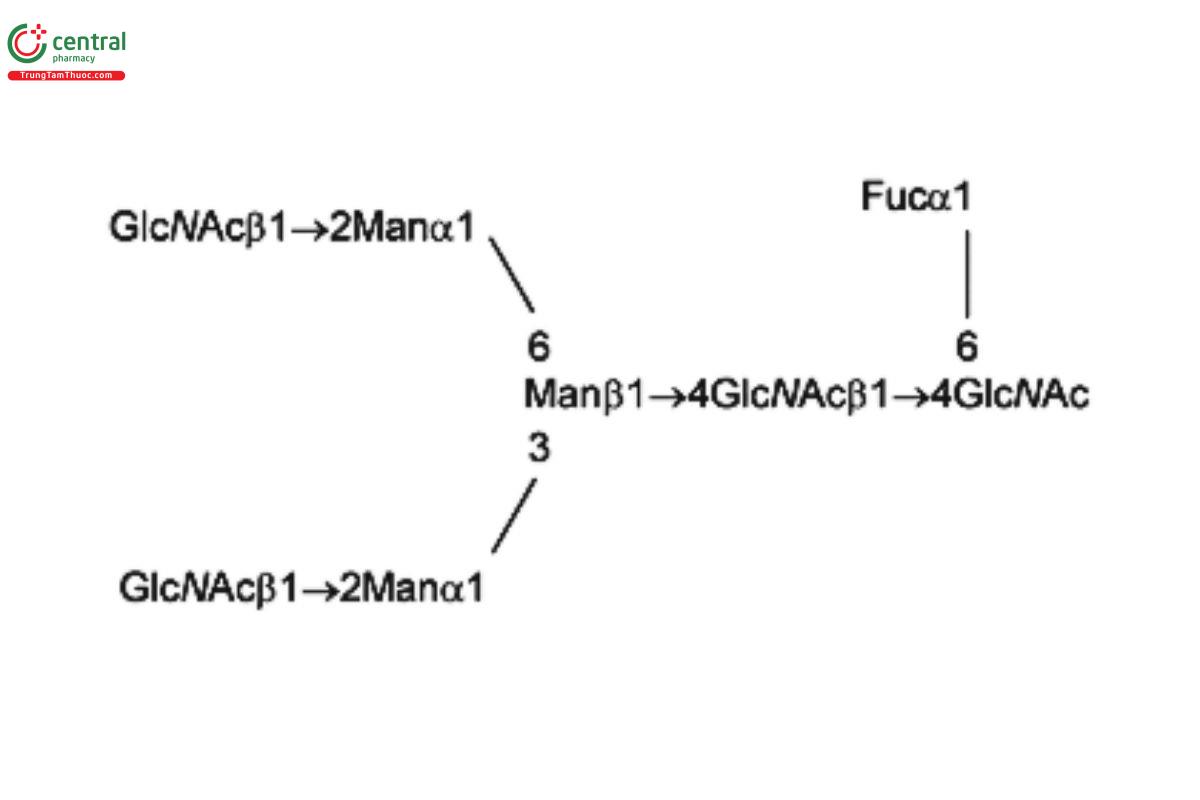
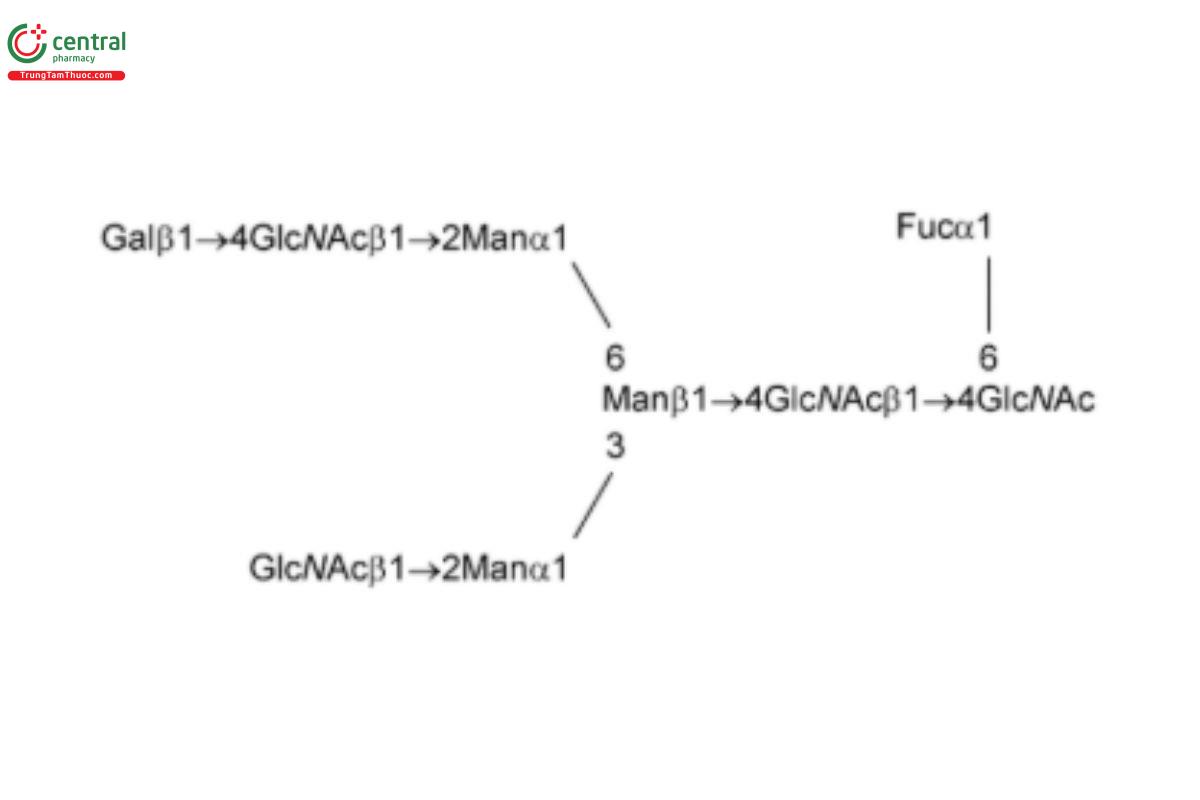
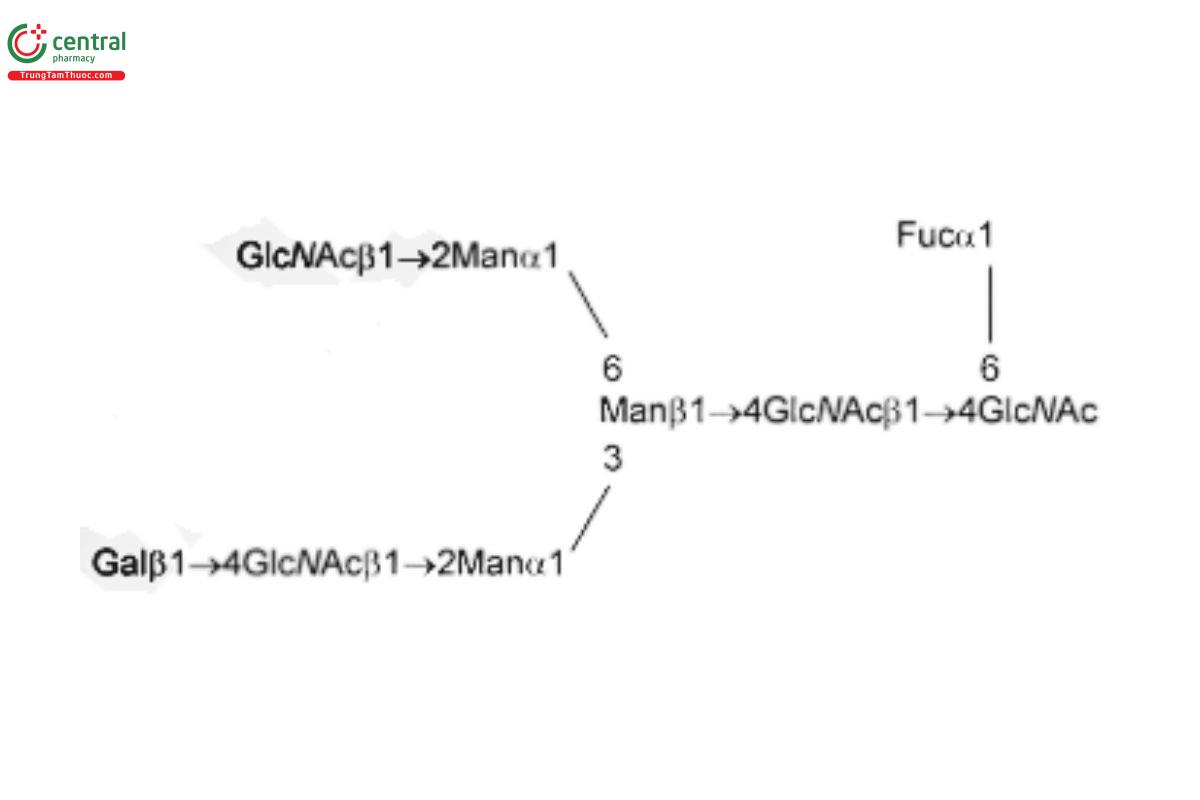
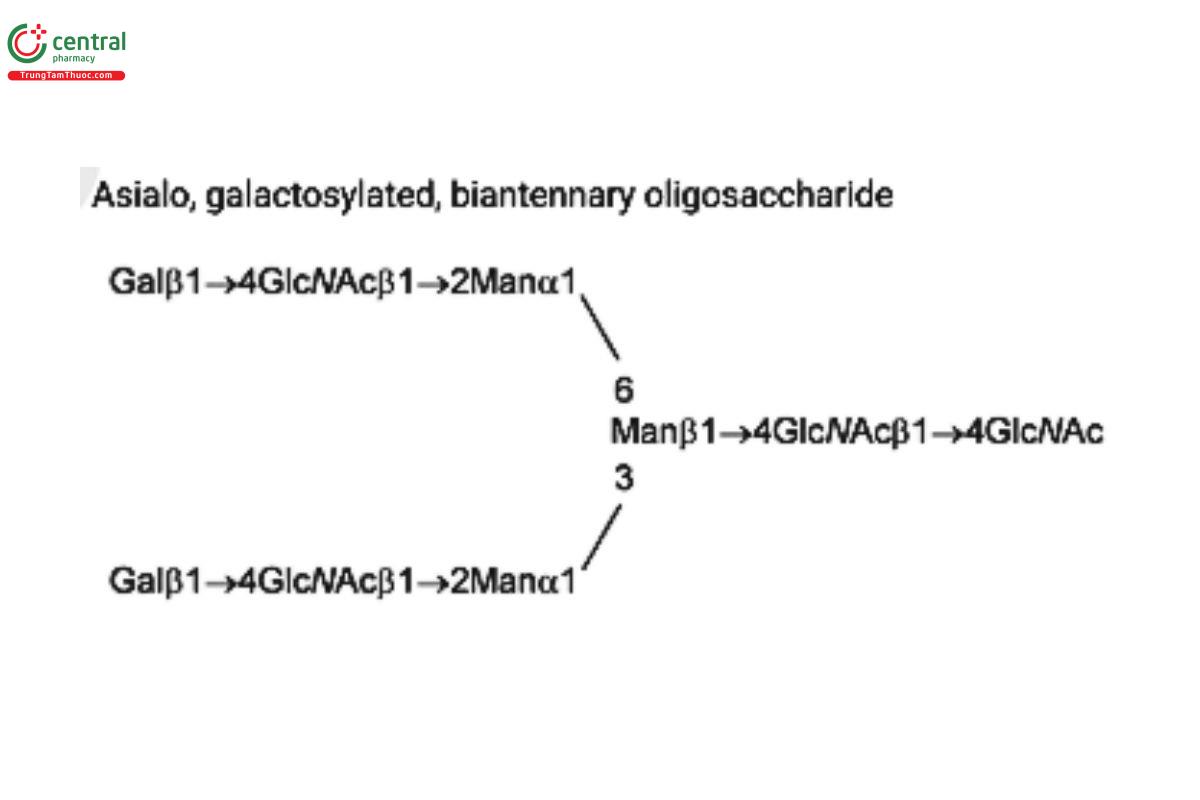
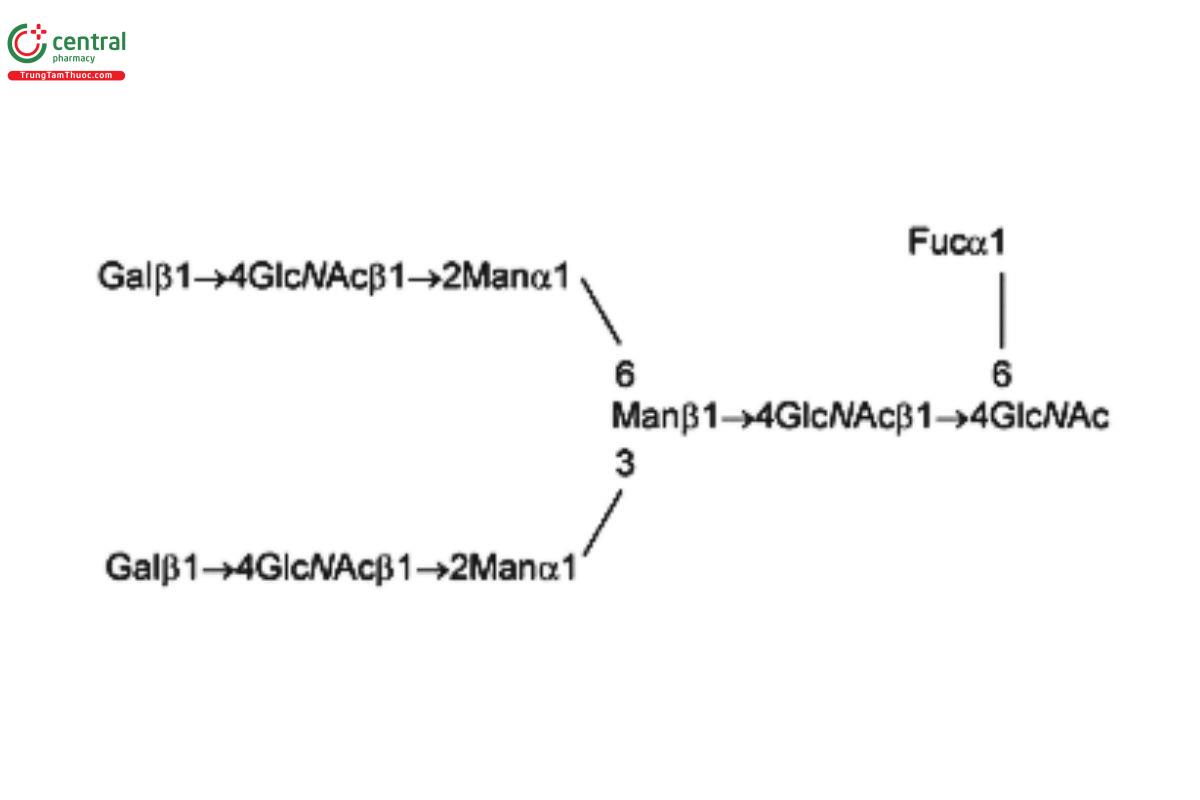
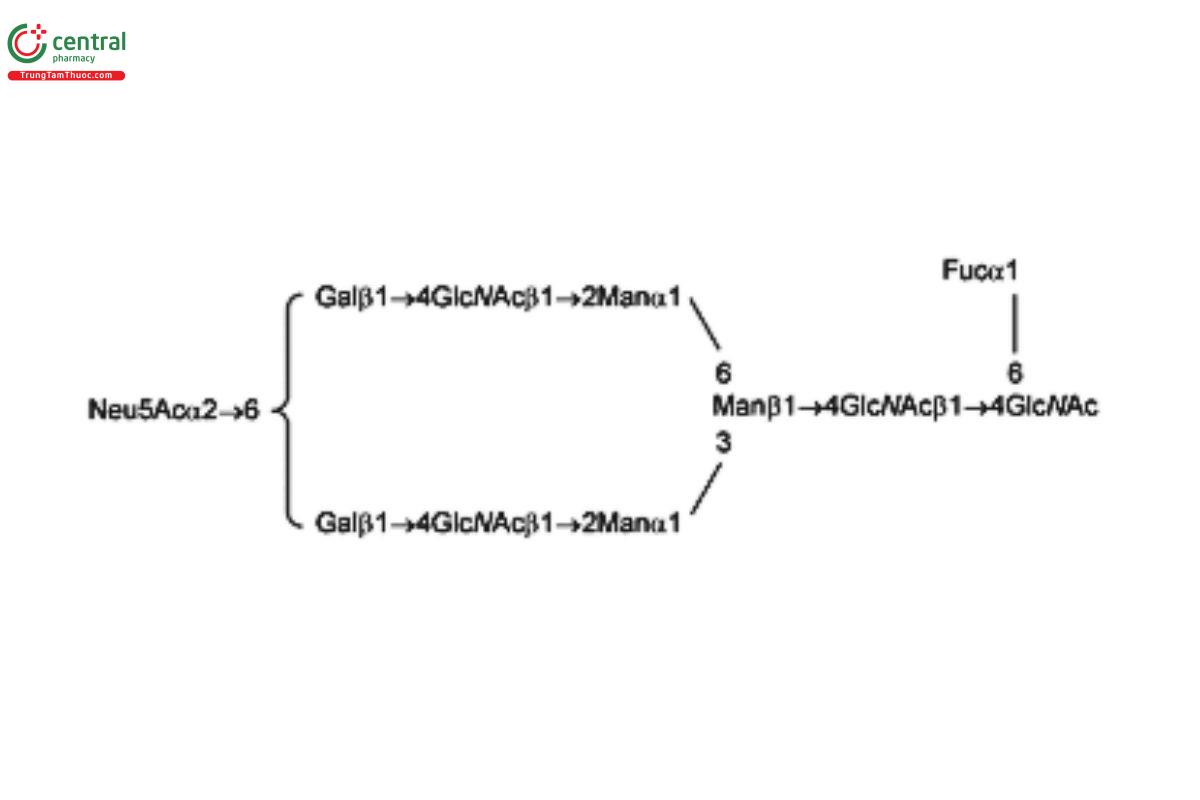
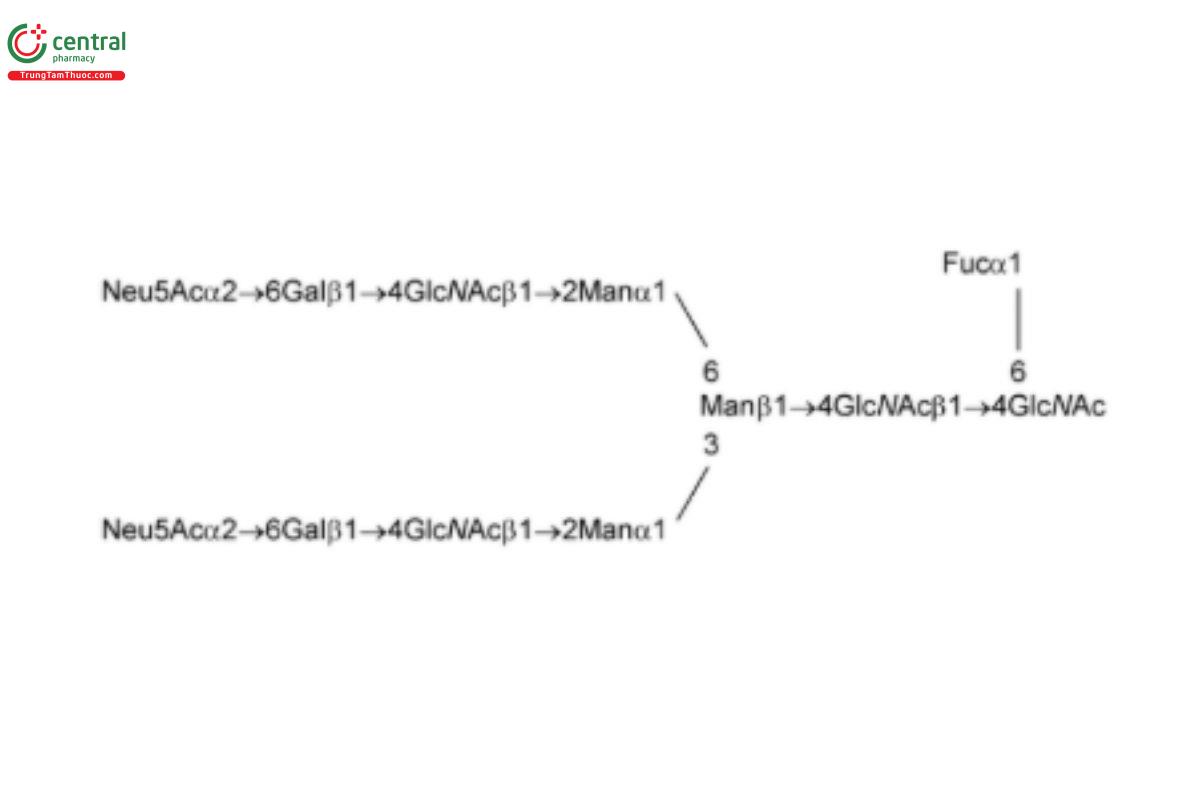
See Appendix (Table 17) for abbreviations and structures of glycans. The sample preparation is for all of the procedures that follow.
Ideally, the samples should be free of salts, excipients, and other carrier materials that may interfere with the analysis. This can be achieved by either 1) dialysis against water, a suitable buffer, or a volatile buffer using an appropriate membrane, 2) sample trapping on a solid-phase extraction (SPE) cartridge followed by washing away of salts and excipients and elution of the required sample, or 3) ultrafiltration using an appropriate membrane.
[Note—In addition to RS, a control sample with a known glycan profile should be included in the overall procedure to confirm correct performance of the analysis. A reaction blank control only containing the buffer matrix of the glycoprotein sample can also be included in the overall procedure.]
• Enzymatic Release of N-Glycans
[Note—The following is a generic method that should be optimized for individual products depending on the amount of glycoprotein to be digested and the glycan structures, especially on the basis of the protein-to-glycan ratio in the molecule and the accessibility of the sugars.]
Procedure 1 for chromatographic separation
Enzyme reaction buffer: 50 mM sodium phosphate pH 7.5 or 15 mM tris(hydroxymethyl)-aminomethane, and adjust with hydrochloric acid to a pH of 7.0.
Digestion with PNGase F: Transfer 0.1–2.25 mg of glycoprotein to a vial, and adjust with Enzyme reaction buffer to a final volume of 30–100 µL. Add PNGase F to the glycoprotein sample in a ratio of 0.5–15 units of PNGase F to 0.1 mg of glycoprotein. Incubate at 37° for 12–24 h. Cool at room temperature for about 5 min. Vortex gently and centrifuge briefly.
[Note—One unit of PNGase F is defined as the amount of enzyme required to catalyze the release of N-linked oligosaccharides from 1.0 nmol of denatured ribonuclease B per min at a pH of 7.5 at 37°, and is equal to 1 IUB milliunit. Complete digestion by PNGase F can be assessed by observing a mobility shift of the deglycosylated proteins on sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), followed by Coomassie Blue staining or on SDS-capillary gel electrophoresis (SDS-CGE), as shown by reduction in mass of approximately 2 kDa per glycan chain cleaved.]
Two methods are described below for performing the separation of released glycans from glycoprotein.
Method 1, using an ultrafiltration membrane with a molecular weight cutoff of 30 kDa:
[Note—Ultrafiltration membranes with a lower molecular weight cutoff may be used for proteins < 150 kDa.]
Remove trace amounts of Glycerin by centrifuging at 11,000 × g and rinsing the membrane with 0.5 mL of water. Discard the permeate. Add 0.1 mL of water into the sample reservoir and then add the digested glycoprotein sample to the reservoir. Rinse the reaction vial with 0.1 mL of water and add to the sample reservoir. Centrifuge at 11,000 × g and collect the permeate. Rinse the sample reservoir twice using 0.1 mL of water for each wash and collect the permeate. Combine the permeates (approximately 0.5 mL) and bring the samples to complete dryness using a centrifugal evaporator without heat.
Method 2, using a reversed-phase SPE cartridge: Add 2.0 mL of methanol into a syringe, attach the syringe to the cartridge, and use the syringe plunger to pass methanol through to waste. Add into a syringe 6.0 mL of 5% (v/v) acetic acid prepared in water. Attach the syringe to the cartridge and use the syringe plunger to pass the acetic acid solution through to waste. Gently apply the digested sample onto an individual cartridge. Into a 1-mL syringe add 0.5 mL of 5% (v/v) acetic acid, attach the syringe to the cartridge, and use the syringe plunger to pass the acetic acid through to waste. To a 2- to 3-mL syringe add 1.5 mL of 5% (v/v) acetic acid. Attach the syringe to the cartridge, and use the syringe plunger to pass the acetic acid solution through, collecting the eluting solution into a 1.5-mL tube. Bring the samples to complete dryness using a centrifugal evaporator without heat.
Procedure 2 for capillary electrophoresis separation
Enzyme reaction buffer: 50 mM sodium phosphate, pH 7.5
Digestion with PNGase F: Add 2 µL of PNGase F (5 units/µL) to 50 µg of glycoprotein sample, and adjust with Enzyme reaction buffer to a final volume of 50 µL. Incubate at 37° for 18 h. Separate released oligosaccharides by centrifugation using a centrifugal filter with a 10,000 molecular weight cut-off. In a centrifugal vacuum evaporator, dry the released oligosaccharides to dryness.
[Note—Follow the guidelines listed in Table 1 to select the appropriate USP RS and label.]
• 2-AB Labeling for Liquid Chromatographic Separation
Solution A: Mix glacial acetic acid and Dimethyl sulfoxide (3:7, v/v).
Solution B: Add 1.5 mL of Solution A to 75 mg of 2-AB. Mix well by gentle vortexing to completely dissolve 2-AB.
Labeling solution: Add 1 mL of Solution B to 63 mg of sodium cyanoborohydride. Mix well by gentle vortexing. Cap the mixture tightly and incubate at 70° for 1–2 min for complete dissolution. Cool at room temperature for 10 min. Use this Labeling solution within 1 h of its preparation, and protect the solution from light.
2-AB labeled standard solution: Add 5–15 µL of Labeling solution to 1 vial of USP Oligosaccharide System Suitability Mixture A RS, USP Oligosaccharide System Suitability Mixture B RS, USP Oligosaccharide System Suitability Mixture C RS, or USP Oligosaccharide System Suitability Mixture D RS, and mix well.
2-AB labeled sample solution: Add 5–15 µL of Labeling solution to the dried glycan sample after enzymatic release and separation steps, and mix well.
Procedure: Immediately incubate the 2-AB labeled standard solution and the 2-AB labeled sample solution at 37° for 16–18 h or at 65° for 2 h. Allow to cool at room temperature for 10 min. Centrifuge briefly.
Dried 2-AB labeled standard or Dried 2-AB labeled sample: Two methods for the removal of free 2-AB are described below.
Method 1, using a gel filtration spin column: Prepare suitable G-10 microcentrifuge spin columns by tapping the column gently to ensure that the dry resin is settled at the bottom. Remove the caps, and place the column in a 2-mL collection tube. Add 0.5 mL of water to the resin, and let it swell for at least 15 min. A longer swelling time (up to 24 h) is acceptable if maintained at 2°–8°. Centrifuge the column at maximum speed for 5–10 s. Remove the water from the collection tube. Wash the resin by adding 0.5 mL of water, and centrifuge the column. Repeat the washing step one more time. After removing the water from the final wash step, spin at maximum speed for 10 s to remove residual water from the resin. Residual water is sufficiently removed if the resin appears white. Place the column in a new, labeled collection tube. Add 100 µL of water to the 2-AB labeled standard solution and the 2-AB labeled sample solution, and then apply the entire solution to the center of each washed G-10 column, respectively. Place the G-10 columns in the microcentrifuge and spin at approximately 200 × g for 1 min. Apply the flow-through (approximately 90–120 µL) to the center of the second unused but previously washed G-10 column and spin at approximately 200 × g for 1 min. Transfer the repeat flow-through (approximately 60–100 µL) to a 0.5-mL microcentrifuge tube. Dry the eluates by centrifugal evaporation with no heat.
Method 2, using an SPE cartridge: Prepare suitable SPE cartridges by washing with 1.0 mL of water, followed by 5 × 1.0 mL of 30% (v/v) acetic acid, and then 1.0 mL of acetonitrile. Apply the 2-AB labeled standard solution and the 2-AB labeled sample solution to the center surface of the cartridge discs, and allow the solution to incubate on the disc for 15–20 min. Wash each disc with 1 mL of acetonitrile, followed by 6 × 1.0 mL of 96% (v/v) acetonitrile, allowing each aliquot to drain before the next is applied. Elute the 2-AB labeled standard solution and the 2-AB labeled sample solution with 3 × 0.5 mL of water, allowing each aliquot to drain before the next is applied. Dry the eluates by centrifugal evaporation with no heat.
• APTS Labeling for Capillary Electrophoresis Separation
APTS labeling reagent: Dissolve 5 mg of trisodium APTS in 48 µL of 15% (v/v) acetic acid.
1 M sodium cyanoborohydride: 1 M sodium cyanoborohydride in tetrahydrofuran
Run buffer: Dissolve 1.492 g of triethanolamine (TEA) and 10 g of glycerol, accurately weighed, in 80 mL of water. Adjust with 1 N hydrochloric acid to a pH of 7.5, and dilute with water to a final volume of 100 mL.
Sample buffer: Dilute 1.0 mL of Run buffer with 9.0 mL of water.
APTS-labeled standard solution: Add 2 µL of the APTS labeling reagent and 2 µL of 1 M sodium cyanoborohydride to a vial of USP Oligosaccharide System Suitability Mixture A RS or USP Oligosaccharide System Suitability Mixture B RS. Incubate at 55° for 90 min. Add 46 µL of water to quench the reaction and mix. Transfer 5 µL of the APTS-labeled USP RS to 1.995 mL of Sample buffer before separation.
APTS-labeled sample solution: Add 2 µL of APTS labeling reagent and 2 µL of 1 M sodium cyanoborohydride to the dried glycan sample after enzymatic release and separation steps. Incubate at 55° for 90 min. Add 46 µL of water to quench the reaction and mix. Transfer 5 µL of the APTS-labeled glycan solution to 95 µL of Sample buffer before separation.
4 SEPARATION AND IDENTIFICATION OF OLIGOSACCHARIDES
• Normal Phase Chromatography/HILIC
Procedure 1
1.4 M formic acid solution: Mix 273 mL of water with 15 mL of 98%–100% formic acid.
1.4 M ammonia solution: Mix 155 mL of water with 40 mL of 26% ammonia solution.
Ammonium formate buffer: Add 1.4 M ammonia solution to 1.4 M formic acid solution until a pH of 4.4 is obtained.
Solution A: Acetonitrile, Ammonium formate buffer, and water (75:4.3:20.7)
Solution B: Acetonitrile, Ammonium formate buffer, and water (54:8.3:37.7)
Mobile phase: See Table 2.
| Time (min) | Solution A (%) | Solution B (%) |
|---|---|---|
| 0.0 | 79 | 21 |
| 80.0 | 47 | 53 |
| 81.0 | 0 | 100 |
| 92.0 | 0 | 100 |
| 93.0 | 79 | 21 |
| 113.0 | 79 | 21 |
Standard solution: Reconstitute the Dried 2-AB labeled standard with NMT 500 µL of water.
Sample solution: Reconstitute the Dried 2-AB labeled sample with an appropriate volume of water.
Blank solution: Buffer matrix of glycoprotein sample carried through the Sample Preparation procedure
Chromatographic system
Mode: LC
Detector: Fluorescence (250-nm excitation wavelength and 428-nm emission wavelength)
Column: 2.0-mm × 15-cm; 3-µm packing L68
Temperatures
Column: 45°
Autosampler: 2°–8°
Flow rate: 0.2 mL/min
Injection volume: 2 µL
System suitability
Samples: Standard solution and Blank solution
Suitability requirements
Blank: No peak in the chromatogram of the Blank solution within the retention time window at 5–113 min
Chromatogram similarity: The chromatogram obtained from the Standard solution corresponds to that of the reference chromatogram provided with the Certificate for either USP Oligosaccharide System Suitability Mixture A RS or USP Oligosaccharide System Suitability Mixture B RS.
Presence of N-glycan species: If USP Oligosaccharide System Suitability Mixture A RS is used, identify peaks corresponding to G0, G0F, G1Fa, G1Fb, G2F, A1F, and A2F in the chromatogram obtained from the Standard solution.
If USP Oligosaccharide System Suitability Mixture B RS is used, identify peaks corresponding to MAN-5, MAN-6, MAN-7, MAN-8, and MAN-9 in the chromatogram obtained from the Standard solution.
Relative retention times: Corresponding to the relative retention times of N-glycans listed in Table 3 or Table 4
Table 3. Standard Solution Using USP Oligosaccharide System Suitability Mixture A RS (HILIC 1)
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| G0 (NGA2) | 0.47 |
| G0F (NGA2F) | 0.57 |
| G1Fa | 0.75 |
| G1Fb | 0.78 |
| G2F (NA2F) | 1.00 |
| A1F (G2fS1) | 1.39 |
| A2F (G2fS2) | 1.76 |
Table 4. Standard Solution Using USP Oligosaccharide System Suitability Mixture B RS (HILIC 1)
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| MAN-5 | 0.74 |
| MAN-6 | 1.00 |
| MAN-7 | 1.29 |
| MAN-8 | 1.60 |
| MAN-9 | 1.85 |
Analysis
Samples: Standard solution and Sample solution
Identify the peaks in the chromatogram obtained from the Sample solution by comparison with the chromatogram obtained from the Standard solution.
Procedure 2
1.4 M formic acid solution: Mix 273 mL of water with 15 mL of 98%–100% formic acid.
1.4 M ammonia solution: Mix 155 mL of water with 40 mL of 26% ammonia solution.
Ammonium formate buffer: Add 1.4 M ammonia solution to 1.4 M formic acid solution until a pH of 4.4 is obtained.
Solution A: Acetonitrile, Ammonium formate buffer, and water (80:4:16)
Solution B: Acetonitrile, Ammonium formate buffer, and water (60:8:32)
Mobile phase: See Table 5.
Table 5.
| Time (min) | Solution A (%) | Solution B (%) |
|---|---|---|
| 0.0 | 20 | 80 |
| 2.0 | 30 | 70 |
| 67.0 | 52 | 48 |
| 67.1 | 80 | 20 |
| 73.0 | 80 | 20 |
| 73.1 | 20 | 80 |
| 85.0 | 20 | 80 |
Standard solution: Reconstitute the Dried 2-AB labeled standard with NLT 500 µL of water. [Note—The volume of water may need to be adjusted so that the uorescence signals from the Standard solution are similar to those from the Sample solution.]
Sample solution: Reconstitute the Dried 2-AB labeled sample with an appropriate volume of water. [Note—Use a ratio of 50 µL of water per 100 µg of glycoprotein used in the Enzymatic Release of N-Glycans test as a starting point.]
Blank solution: Buffer matrix of glycoprotein sample carried through the Sample Preparation procedure
Chromatographic system
(See Chromatography 〈621〉, System Suitability.)
Mode: LC
Detector: Fluorescence; set the excitation and emission wavelengths as listed in Table 6
| Time (min) | Excitation (nm) | Emission (nm) |
|---|---|---|
| 0.00 | 330 | 420 |
| 1.00 | 330 | 420 |
| 1.01 | 400 | 500 |
| 10.00 | 400 | 500 |
| 10.01 | 330 | 420 |
| 85.00 | 330 | 420 |
Column: 4.6-mm × 25-cm; 5-µm packing L68
Temperatures
Column: 35°
Autosampler: 2°–8°
Flow rate: 0.4 mL/min
Injection volume: 10 µL
System suitability
Sample: Standard solution
Suitability requirements
Chromatogram similarity: The chromatogram obtained from the Standard solution corresponds to that of the reference
chromatogram provided with the Certicate for either USP Oligosaccharide System Suitability Mixture A RS or USP Oligosaccharide
System Suitability Mixture B RS.
Presence of N-glycan species: If USP Oligosaccharide System Suitability Mixture A RS is used, identify peaks corresponding to G0, G0F, G1Fa, G1Fb, G2, G2F, A1F, and A2F (see Table 7; see also Appendix) in the chromatogram obtained from the Standard solution by comparing the reference chromatogram provided with the USP Certicate for USP Oligosaccharide System Suitability Mixture A RS.
If USP Oligosaccharide System Suitability Mixture B RS is used, identify peaks corresponding to MAN-5, MAN-6, MAN-7, MAN-8, and MAN-9 (see Table 8; see also Appendix) in the chromatogram obtained from the Standard solution by comparing the reference chromatogram provided with the USP Certicate for USP Oligosaccharide System Suitability Mixture B RS. Relative retention times: Corresponding to the relative retention times of N-glycans listed in Table 7 or Table 8
Table 7. Standard Solution Using USP Oligosaccharide System Suitability Mixture A RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| G0 (NGA2) | 0.78 |
| G0F (NGA2F) | 0.83 |
| G1Fa | 0.91 |
| G1Fb | 0.92 |
| G2 (NA2) | 0.96 |
| G2F (NA2F) | 1.00 |
| A1F (G2fS1) | 1.11 |
| A2F (G2fS2) | 1.20 |
Table 8. Standard Solution Using USP Oligosaccharide System Suitability Mixture B RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| MAN-5 | 0.77 |
| MAN-6 | 0.86 |
| MAN-7 | 0.93 |
| MAN-8 | 1.00 |
| MAN-9 | 1.05 |
Analysis
Sample: Sample solution
Integrate peaks in the resulting chromatogram and report the relative peak areas of glycan structures relevant to the product. For general information on chromatographic peak integration, see 〈621〉.
High-Performance Anion Exchange Liquid Chromatography with Fluorometric Detection
Procedure 1
Solution A: Water; degas before use.
Solution B: 0.5 M sodium acetate, prepared as follows. Dissolve 20.5 g of anhydrous sodium acetate in 450 mL of water and mix well.
Dilute with water to final volume of 500 mL. Filter the solution through a membrane of NMT 0.45-μm pore size, and degas before use.
Solution C: 0.5 M sodium acetate, prepared as follows. To 972 mL of water, add 26 mL of 50% (w/w) sodium hydroxide solution. Pass the solution through an alkaline-resistant
lter membrane of NMT 0.45-μm pore size, and degas before use.
Mobile phase: See Table 9.
| Time (min) | Solution A (%) | Solution B (%) | Solution C (%) | Elution |
|---|---|---|---|---|
| 0.0 | 70 | 0 | 30 | Initial condition |
| 15.0 | 70 | 0 | 30 | 0–75 mM sodium acetate |
| 65.0 | 55 | 15 | 30 | |
| 66.0 | 0 | 50 | 50 | 250 mM sodium acetate wash |
| 74.0 | 0 | 50 | 50 | |
| 75.0 | 70 | 0 | 30 | Re-equilibrium |
| 90.0 | 70 | 0 | 30 |
Standard solution: Reconstitute the Dried 2-AB labeled standard with NMT 500 µL of water. [Note—The volume of water may need to be adjusted so that the fluorescence signals from the Standard solution are similar to those from the Sample solution.]
Sample solution: Reconstitute the Dried 2-AB labeled sample with an appropriate volume of water. [Note—Use a ratio of 20 µL of water per 100 µg of glycoprotein used in the Enzymatic Release of N-Glycans test as a starting point.]
Blank solution: Buffer matrix of glycoprotein sample carried through the Sample Preparation procedure
Chromatographic system
(See Chromatography 〈621〉, System Suitability.)
Mode: LC
Detector: Fluorescence (330-nm excitation wavelength and 420-nm emission wavelength)
Columns
Guard: 4.0-mm × 5-cm; 10-µm packing L46
Analytical: 4.0-mm × 25-cm; 10-µm packing L46
Temperatures
Column: 25°
Autosampler: 4°
Flow rate: 0.8 mL/min
Injection volume: 50 µL
System suitability
Sample: Standard solution
Suitability requirements
Chromatogram similarity: The chromatogram obtained from the Standard solution corresponds to that of the reference chromatogram provided with the Certificate for either USP Oligosaccharide System Suitability Mixture A RS or USP Oligosaccharide System Suitability Mixture B RS.
Presence of N-glycan species: If USP Oligosaccharide System Suitability Mixture A RS is used, identify peaks corresponding to G0F, G1F, G0, G2F, A1F, and A2F (see Table 10; see also Appendix) in the chromatogram obtained from the Standard solution by comparing the reference chromatogram provided with the USP Certificate for USP Oligosaccharide System Suitability Mixture A RS. If USP Oligosaccharide System Suitability Mixture B RS is used, identify peaks corresponding to MAN-5, MAN-6, MAN-7, MAN-8, and MAN-9 (see Table 11; see also Appendix) in the chromatogram obtained from the Standard solution by comparing the reference chromatogram provided with the USP Certificate for USP Oligosaccharide System Suitability Mixture B RS.
Relative retention times: Corresponding to the relative retention times of N-glycans listed in Table 10 or Table 11
Table 10. Standard Solution Using USP Oligosaccharide System Suitability Mixture A RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| G0F (NGA2F) | 0.67 |
| G1F | 0.86 |
| G0 | 0.92 |
| G2F | 1.00 |
| A1F | 1.75 |
| A2F | 2.22 |
Table 11. Standard Solution Using USP Oligosaccharide System Suitability Mixture B RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| MAN-5 | 0.86 |
| MAN-6 | 1.00 |
| MAN-7 | 1.09 |
| MAN-8 | 1.15 |
| MAN-9 | 1.24 |
Analysis: Equilibrate the Chromatographic system with the initial Mobile phase conditions described in Table 9 for a minimum of 15 min. Inject 50 µL of water for the first injection.
Sample: Sample solution
Integrate peaks in the resulting chromatogram and report the relative peak areas of glycan structures relevant to the product. For general information on chromatographic peak integration, see 〈621〉.
Procedure 2
Solution A: Water; degas before use
Solution B: 0.5 M sodium acetate, prepared as follows. Dissolve 41.0 g of sodium acetate anhydrous in 900 mL of water. Transfer the solution to a 1-L volumetric flask, and bring with water to volume. Filter the solution through a nylon membrane with NMT 0.45-µm pore size, and degas before use.
Solution C: 0.5 M sodium hydroxide, prepared as follows. To 900 mL of water, add 26 mL of 50% (w/w) sodium hydroxide solution, and mix well. Bring the solution with water to the final volume of 1000 mL. Filter the solution through an alkaline-resistant nylon membrane with NMT 0.45-µm pore size, and degas before use.
Mobile phase: See Table 12.
| Time (min) | Solution A (%) | Solution B (%) | Solution C (%) | Elution |
|---|---|---|---|---|
| 0.0 | 80 | 10 | 10 | Initial condition |
| 15.0 | 80 | 10 | 10 | 50–150 mM sodium acetate |
| 70.0 | 60 | 30 | 10 | 150–450 mM sodium acetate |
| 94.0 | 0 | 90 | 10 | Sodium acetate wash |
| 105 | 0 | 10 | 90 | Sodium hydroxide wash |
| 111 | 80 | 10 | 10 | |
| 130 | 80 | 10 | 10 | Re-equilibrium |
Standard solution: Reconstitute the Dried 2-AB labeled standard with NMT 500 µL of water.
Sample solution: Reconstitute the Dried 2-AB labeled sample with an appropriate volume of water.
Blank solution: Buffer matrix of glycoprotein sample carried through the Sample Preparation procedure
Chromatographic system
(See Chromatography 〈621〉, System Suitability.)
Mode: LC
Detector: Fluorescence (330-nm excitation wavelength and 420-nm emission wavelength)
Columns
Guard: 4.0-mm × 5-cm; 10-µm packing L46
Analytical: 4.0-mm × 25-cm; 10-µm packing L46
Temperatures
Column: 25°
Autosampler: 4°
Flow rate: 0.8 mL/min
Injection volume: 50 µL
System suitability
Sample: Standard solution
Suitability requirements
Chromatogram similarity: The chromatogram obtained from the Standard solution corresponds to that of the reference chromatogram provided with the Certificate for either USP Oligosaccharide System Suitability Mixture C RS or USP Oligosaccharide System Suitability Mixture D RS.
Presence of N-glycan species: If USP Oligosaccharide System Suitability Mixture C RS is used, identify peaks corresponding to bi-, tri-, and tetra-antennary N-linked oligosaccharides with variable sialylation. If USP Oligosaccharide System Suitability Mixture D RS is used, identify peaks corresponding to sialylated bi- and tri-antennary complex type structures with an additional sialic acid on one of the tri-antennary structures.
Relative retention times: Corresponding to the relative retention times of N-glycans listed in Table 13 or Table 14
Table 13. Standard Solution Using USP Oligosaccharide System Suitability Mixture C RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| A1 (G1S1), peak 1 | 0.31 |
| A1 (G1S1), peak 2 | 0.32 |
| A2 (G2S2) | 0.73 |
| A3 (G3S3) | 1.00 |
Table 14. Standard Solution Using USP Oligosaccharide System Suitability Mixture D RS
| N-Glycan | Approximate Relative Retention Time |
|---|---|
| A1 (G1S1), peak 1 | 0.23 |
| A2 (G2S2) | 0.57 |
| A3 (G3S3) | 0.78 |
| A3G3S4 | 1.00 |
Analysis: [NOTE—Equilibrate the column with initial mobile phase conditions for a minimum of 15 min.]
Sample: Sample solution
Inject 25 µL of water, and run the gradient program at least once to equilibrate the column and system. Integrate peaks in the resulting chromatogram, and report the relative peak areas of glycan structures relevant to the product. For general information on chromatographic peak integration, see 〈621〉.
5 CAPILLARY ELECTROPHORESIS
Run buffer: Dissolve 1.492 g of TEA and 10 g of Glycerol in 80 mL of water. Adjust with 1 N hydrochloric acid to a pH of 7.5 and dilute with water to a final volume of 100 mL.
Standard solution: APTS-labeled standard solution prepared as directed in the test for APTS Labeling for CE Separation
Sample solution: APTS-labeled sample solution prepared as directed in the test for APTS Labeling for CE Separation
Blank solution: Buffer matrix of glycoprotein sample carried through the Sample Preparation procedure
Instrumental conditions
Mode: CE
Detector: Laser-induced fluorescence (488-nm excitation wavelength and 520-nm emission wavelength)
Capillary: 50-µm inner diameter bare fused-silica with 50 cm total length and 40 cm separation length
Preconditioning of the capillary: Between each run, rinse for 5 min at 40 psi with 0.5 N sodium hydroxide, followed by water for 1 min at 40 psi. [Note—Precondition a new capillary or as needed: Rinse for 5 min at 20 psi with methanol, followed by water for 1 min at 50 psi, 1 N hydrochloric acid for 5 min at 20 psi, water for 1 min at 50 psi, 0.5 N sodium hydroxide for 25 min at 20 psi, and water for 5 min at 50 psi.]
Prefilling of the capillary: Rinse for 5 min at 40 psi with Run buffer.
Sample injection: 10 s hydrodynamic (pressure) injection
Voltage: 22 kV for 60 min
Temperatures
Capillary cartridge: 18°
Sample storage: 20°
System suitability
Sample: Standard solution
Suitability requirements
Electropherogram similarity: The electropherogram obtained from the Standard solution corresponds to that of the reference electropherogram provided with the Certificate for either USP Oligosaccharide System Suitability Mixture A RS or USP Oligosaccharide System Suitability Mixture B RS.
Presence of N-glycan species: If USP Oligosaccharide System Suitability Mixture A RS is used, identify peaks corresponding to G2F, G2, G1Fa, G1Fb, G0F, MAN-5, G0, and A1F (see Table 15; see also Appendix) in the electropherogram obtained from the Standard solution by comparing the reference electropherogram provided with the USP Certificate for USP Oligosaccharide System Suitability Mixture A RS. If USP Oligosaccharide System Suitability Mixture B RS is used, identify peaks corresponding to MAN-5, MAN-6, MAN-7, MAN-8, and MAN-9 (see Table 16; see also Appendix) in the electropherogram obtained from the Standard solution by comparing the reference electropherogram provided with the USP Certificate for USP Oligosaccharide System Suitability Mixture B RS.
Relative migration times: Corresponding to the relative migration times listed in Table 15 or Table 16
Table 15
| N-Glycan | Approximate Relative Migration Time |
|---|---|
| G2F | 0.79 |
| G2 | 0.84 |
| G1Fb | 0.87 |
| G1Fa | 0.90 |
| G0F | 1.00 |
| MAN-5 | 1.03 |
| G0 | 1.12 |
| A1F | 1.18 |
Table 16
| N-Glycan | Approximate Relative Migration Time |
|---|---|
| MAN-9 | 0.82 |
| MAN-8 | 0.85 |
| MAN-7 | 0.91, 0.92, 0.94 |
| MAN-6 | 1.00 |
| MAN-5 | 1.11 |
Analysis
Sample: Sample solution
Integrate peaks in the resultant electropherogram and report the relative peak areas of glycan structures relevant to the product.
6 ADDITIONAL REQUIREMENTS
USP Reference Standards 〈11〉
USP Oligosaccharide System Suitability Mixture A RS
USP Oligosaccharide System Suitability Mixture B RS
USP Oligosaccharide System Suitability Mixture C RS
USP Oligosaccharide System Suitability Mixture D RS
Table 17
| Glycan | Description |
|---|---|
| G0 (NGA2) | Asialo-, agalacto- biantennary oligosaccharide
|
| G0F (NGA2F) | Asialo-agalacto-fucosylated biantennary oligosaccharide
|
| G1Fa (NA1F, FA2G1, or NA2G1F) with the α(1,6) linkage | Asialo-, monogalactosylated, fucosylated biantennary oligosaccharide with galactosylation at the α(1,6) linkage
|
| G1Fb (NA1F, FA2G1, or NA2G1F) with the α(1,3) linkage | Asialo-, monogalactosylated, fucosylated biantennary oligosaccharide with galactosylation at the α(1,3) linkage
|
| G2 (NA2) | Asialo, galactosylated, biantennary oligosaccharide
|
| G2F (NA2F) | Asialo-fucosylated biantennary oligosaccharide
|
| A1F (G2fS1) | Monosialo-fucosylated biantennary oligosaccharide
|
| A2F (G2fS2) | Disialo-fucosylated biantennary oligosaccharide
|
| MAN-5 | Oligomannose 5; Oligomannose N-linked oligosaccharide with 5 mannosyl residues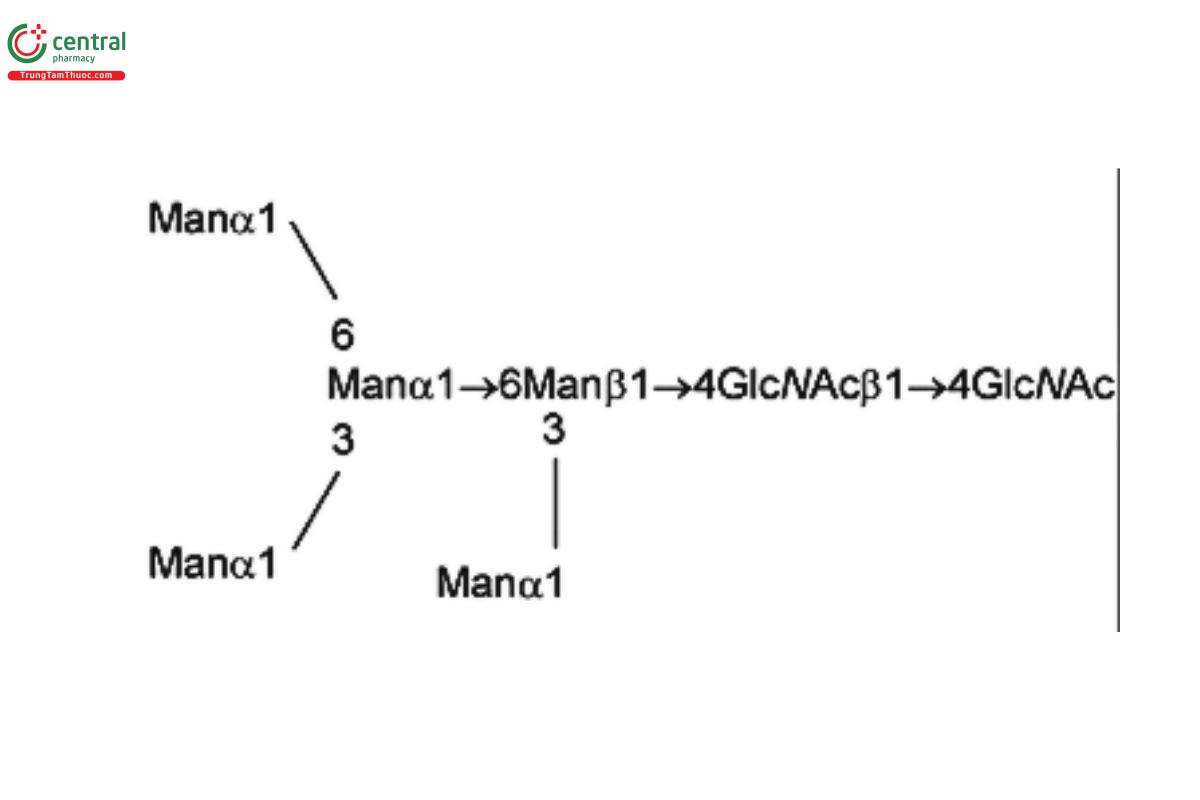 |
| MAN-6 | Oligomannose 6; Oligomannose N-linked oligosaccharide with 6 mannosyl residues
|
| MAN-7 | Oligomannose 7; Oligomannose N-linked oligosaccharide with 7 mannosyl residues
|
| MAN-8 | Oligomannose 8; Oligomannose N-linked oligosaccharide with 8 mannosyl residues
|
| MAN-9 | Oligomannose 9; Oligomannose N-linked oligosaccharide with 9 mannosyl residues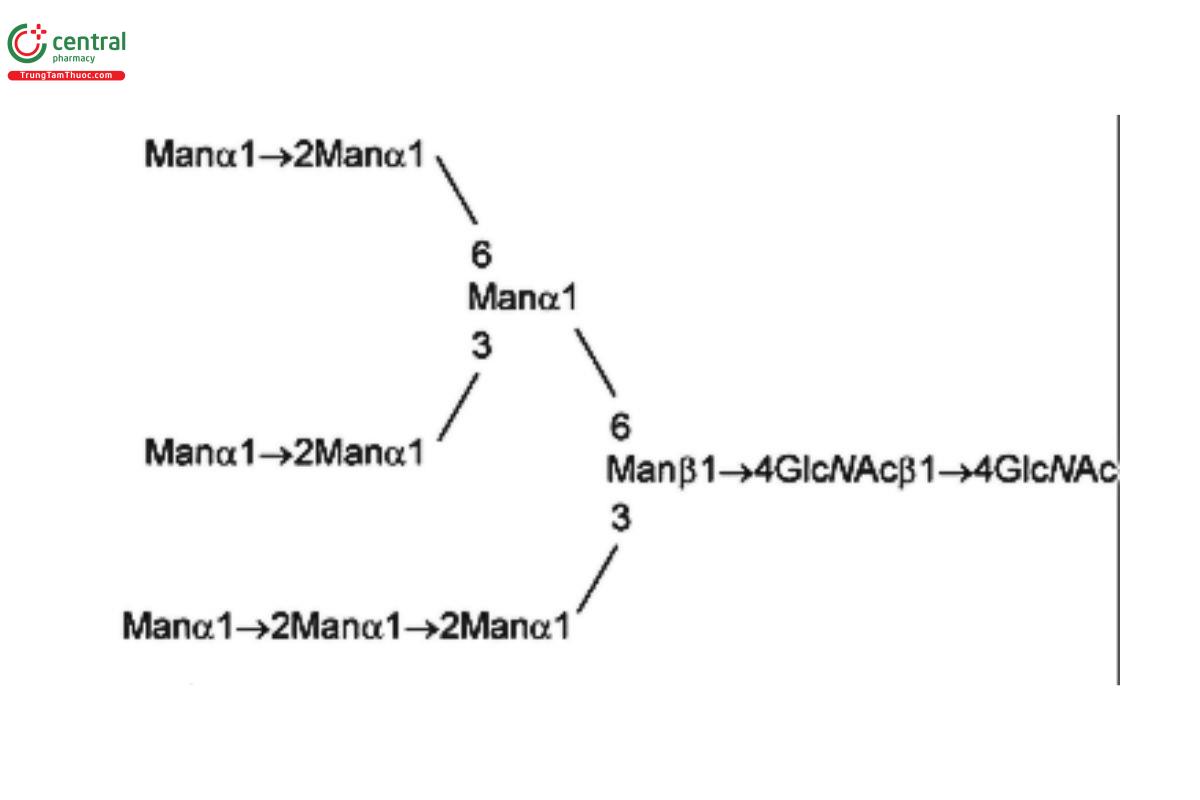 |
| A1 (G1S1) | Mono-sialylated, galactosylated, biantennary oligosaccharide
|
| A2 (G2S2) | Di-sialylated, galactosylated, biantennary oligosaccharide
|
| A3 (G3S3) | Tri-sialylated, galactosylated, triantennary oligosaccharide
|
| A3G3S4 (S4NA3, or A3 + Sa) | Tetra-sialylated, galactosylated, triantennary oligosaccharide
|
1 A suitable ultrafiltration membrane is YM-30 microcon centrifugal filter device from Millipore (catalog numbers 42422, 42409, and 42410) or an equivalent.
2 A suitable ultrafiltration membrane is EMD Millipore Ultra-0.5 Centrifugal Filter Unit (catalog number UFC501096) or an equivalent.
3 A suitable SPE cartridge is Glyko GlycoClean R Cartridge from Prozyme (catalog number GKI-4756) or an equivalent.
4 A suitable 2-AB labeling kit is Glyko Signal 2-AB Labeling Kit from Prozyme (catalog number GKK-404), LudgerTag 2-AB labeling kit from QA Bio (catalog number LT-KAB-A2), or an equivalent.
5 A suitable prepacked Sephadex G-10 column is Macro Spin G-10 mini SEC column from Harvard Apparatus (catalog number 743900) or The Nest Group (catalog number SMM S010), or an equivalent.
6 A suitable cartridge to remove free 2-AB is Glyko GlycoClean S Cartridge from Prozyme (catalog number GKI-4726) or an equivalent.